|
You will write the Results and Discussion for your enzyme inhibition study. Additionally, you will present your calculations in Excel. |
Excel Spreadsheet
Highlight the cells containing specific sample calculations by filling those cells in yellow. The purpose of this is to help the grader find the required information. Neglecting to do this will result in points deductions.
- Create plots of A400 vs. time for all your raw kinetics data for the eight catechol concentrations with and without inhibitor. Graph trendlines on each data set to illustrate how well each approximates linear intitial rates. Use these plots to guide any decisions about elimination of later time points. You are not required to label the axes.
- Clearly label your substrate concentration calculations (the final concentrations in the assay) and highlight the cell of one sample calculation in yellow.
- Clearly label the slope calculations and highlight the cell of one sample calculation in yellow.
- Clearly label the relative activity calculations and highlight one sample calculation in yellow.
- Create a table that summarizes your [catechol] and vo (relative activity) data.
- Clearly label the values for your Eadie-Hofstee plots and highlight one sample calculation for the x-axis calculation in yellow.
- Create Eadie-Hofstee plots and graph a linear trendline of the data. These graphs will be used in your Results.
- In the LINEST array of your Eadie-Hofstee plot data, the four cells for slope, slope standard error, y-intercept, and y-intercept standard error must be highlighted in yellow.
- Clearly label the cells containing calculations and error propagation for Vmax/Km and highlight these cells in yellow.
- Create an organized summary table of your estimates for Vmax, Vmax/Km, and Km with estimated uncertainty. f you organize these data into a formal table, you can simply cut and paste it into your writing assignment as Table 1 (see details below).
- Calculate the inhibitor concentration and use this to calculate your value(s) for Ki in μM units. Clearly label and highlight these cells in yellow .
- Clearly label any equation(s) used to propagate error for the Ki value(s). Label and highlight these cells in yellow.
Results and Discussion
Figure 1. This figure will present representative data that demonstrates the quality of your kinetics measurements. It is the common to only provide a plot of one dataset, either the uninhibited data or the inhibitied data if the data quality is comparable. However, if one time-course dataset has significant problems, ethically, you must report the lower-quality dataset. It is acceptable to re-use Figure 1 of Assignment 4 if those data are representative. As a reminder, a good caption should identify the enzyme, substrate, inhibitor, buffer, and temperature (see the "Enzyme Kinetics Data" entry in the Methods-Captions tab of the Writing Guidelines ). No equations are necessary in the caption of this figure.
Figure 2. Prepare an Eadie-Hofstee plot of the kinetics data from Table 1, make sure to include both labels and the correct units on the axes. Plot a linear trendline through your data, include equation and R2 in the caption. Your caption should also name the enzyme, temperature, buffer conditions, the substrate, the inhibitor and its concentration with the correct units, the values of your inhibition constant(s), and inhibtion mechanism.
Table 1. Create a table summarizing your estimates for Vmax, Vmax/Km, and Km for tyrosinase reaction of catechol at pH 6.5 both with and without inhibitor. Include uncertainty for all six parameters. Identify the inhibitor and its concentration.
State your objective.
- Suggest an uresolved problem in the current literature that you will address then state the objective of your investigation and describe how it will help to resolve that problem.
- To help you justify the purpose of this study (there is no one correct justification), it is helpful for you to read the introductions of journal articles that report Ki values for your inhibitor. One such paper (Chiari 2011) is posted with this week's assignment on the D2L main page. From such papers, you will learn why other researchers consider tyrosinase enzyme worthy of study.
- To relate your study to other work in the field, consider your data to be a unique contribution to the current state of tyrosinase research.
- After providing general background, raise an issue that has not yet been studied in the published literature. Some ideas to consider:
- Has the Ki been reported for your inhibitor in Agaricus bispous tyrosinase? If not, your measurement is novel and can be compared to other known Ki values for your inhibitor.
- You studied the inhibition mechanism of your tyrosinase inhibitor with SDS activation. Have any tyrosinase inhibition studies been conducted with SDS activation? Would you predict that SDS changes the mechanism? Does it change the mechanism?
After you state your objectives, briefly describe the experimental design of work you will present.
- When you describe the experiment, you must mention the source organism of tyrosinase and origin of the tyrosinase that you studied (i.e. a commercial sample, an extract from a specific tissue, etc.).
Present your results
- Write a narrative that explains the key results of each figure and table.
- Make in-text references to every figure and table.
- Comment on the quality of your data (the quality of the trends (R2), the number of data points, the RSDs for your kinetics parmeters, etc.).
- If you do not trust your results, explain what went wrong.
- If you dropped data points, justify your decisions.
Discussion
- Analyze the statistical significance of the change in your relevant kinetics parameters with and without inhibitor in Table 2.
- Interpret this analysis to determine the mechanism of your inhibitor.
- Compare your Ki value(s) and mechanism to a literature value for tyrosinase inhibition constants reported for the same inhibitor.
- If BRENDA does not list Ki values your inhibitor in Agaricus bisporus, you should compare your values to tyrosinase in other organisms.
- Compare your catechol Km to published catechol Km values with tyrosinase in Agaricus bisporus.
- If your Km differs significantly from the literature, suggest a reason for why that might be the case.
- Include at least one literature citation in the appropriate format for each published reference value.
- As you interpret your results, be clear about the limits of your ability to interpret based on the magnitude of your uncertainty.
- Summarize whether or not you were successful in achieving your goals.
- Suggest at least one future experiment or improvement.
Submit
- A printed copy of the Lab 6 Assignment with the rubric stapled on top.
- Upload to the D2L Submissions folder:
- A digital copy of the Lab 6 Assignment
- Your Excel Spreadsheet presented according to the instructions above
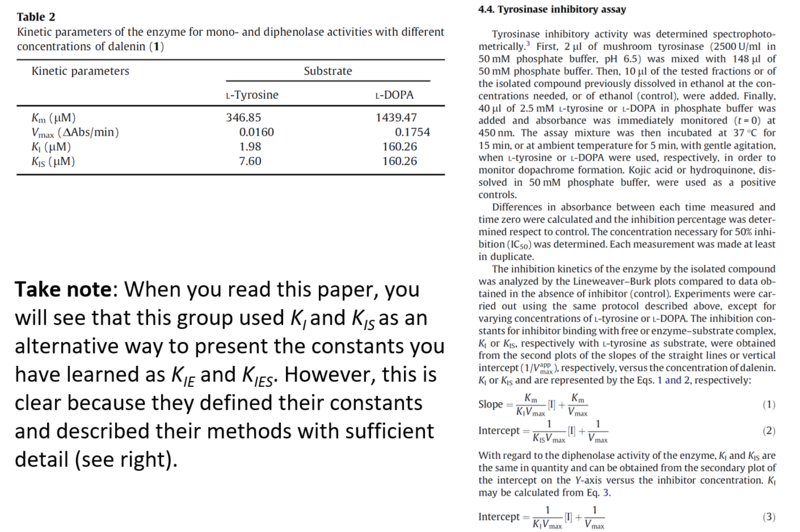
Appendix: An Example of a Published Tyrosinase Inhibition Report
(The full text is available on D2L as "Chiari 2011")
Video Lecture:

You can find morehelpful articcles like this by going to BRENDA, searching "tyrosinase", selecting "Ki" under functional parameters, and selecting papers for Agaricus bisporus.

The figures in this paper illustrate a clear and complete presentation of enzyme inhibition data. Even though this article uses the Lineweaver-Burk method of displaying their data, their clear presentation allows the reader to be aware of this, and to take the data with a grain of salt.